Inflammasome is a central player in the induction of obesity and insulin resistance. Pathway analysis of expression data: deciphering functional building blocks of complex diseases. Isserlin R, Merico D, Alikhani-Koupaei R, Gramolini A, Bader GD, Emili A. Pathway analysis of dilated cardiomyopathy using global proteomic profiling and enrichment maps Nature Blogs - In the news (The Globe and Mail) Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R, Conroy J, Magalhaes TR, Correia C, Abrahams BS, Almeida J, Bacchelli E, Bader GD, et al. Merico D, Isserlin R, Stueker O, Emili A, Bader GDįunctional impact of global rare copy number variation in autism spectrum disorders. How to convert DAVID gene-sets to GMT: R scriptĮnrichment Map: A Network-Based Method for Gene-Set Enrichment Visualization and Interpretation GSEA) by querying public resources such as Gene Ontology and KEGG, and using Entrez-Gene ID for genes we generate gene-set files for enrichment analysis (e.g.Description of sources and methods used to create collection can be found here User Guide for older 2.2 version of EnrichmentMap (including tutorials) is here: ĭirectly downloading our monthly updated gene-set collections from Baderlab genesets collections. The EnrichmentMap User Guide has been moved to: Tutorials for the older 2.2 version of EnrichmentMap are here: Tuturials and protocols for EnrichmentMap 3.0 are here: This will install EnrichmentMap as well as a suite of other apps that work well with EnrichmentMap, including AutoAnnotate, WordCloud and clusterMaker2: Download and customize legend to fit your networkĮnrichmentMap is available on the Cytoscape App Store: Īlternatively download the EnrichmentMap Pipeline Collection. Enrichment Map also enables the comparison of two different enrichment results in the same map.įollow this link for more details and an analysis example.įor easier figure creation download our template legends that contains all the visual properties used in a standard enrichment map. In this way, mutually overlapping gene-sets cluster together, making interpretation easier. Gene-sets, such as pathways and Gene Ontology terms, are organized into a network (i.e. Enrichment results have to be generated outside Enrichment Map, using any of the available methods. Cytoscape Editor and Metanodes PlugIn do not interact well.Enrichment Map is a Cytoscape plugin for functional enrichment visualization.Reimplementation of groups and metanodes is planned for Cytoscape 2.5 Metanodes classes in the library will change in the future. IMPORTANT NOTE: Implementation of MetaNodeUtils class, and other You include it in your plugIn's jar file. Want to load a plugIn into Cytoscape that makes use of it, make sure To use the library you can include its jar in your classpath. The Metanodes Library is a Java package for Cytoscape that containsĬlasses and methods to create, collapse, expand, and destroy metanodes. MetaNode commands are now available as node context menus or under the plugins menu.Copy metanodepl.jar to the plugins folder of your Cytoscape installation.Metanode by doing a performing operation on its children nodes. The PlugIn transfers node attributes to the new That are composed of metanodes, thus creating a hierarchical structure 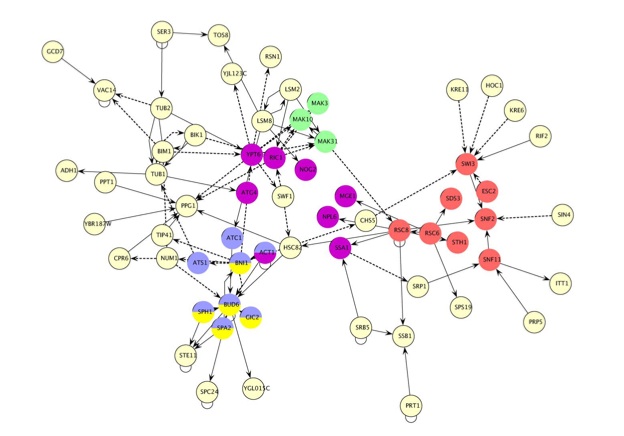
The Metanodes PlugIn is a Cytoscape PlugIn that provides a dialog (seeįigure below) that allows users to select a group of nodes in a networkĪnd create a metanode that represents them. Metanodes simplify complex networks and can be used to represent complex hierarchical systems in two dimensions. A metanode at an even higher level of abstraction can represent a group of protein complexes (metanodes can be recursive). For example, a metanode can represent a protein complex if it's subgraph is composed of individual nodes and edges representing interacting proteins. To go meta means to take a step back and look at the bigger picture.Ī metanode is a graph node that represents a subgraph that is at a lower level of abstraction. Meta: Something on an abstraction level higher than the current. Previously at the Hood Lab, Institute for Systems Biology, Seattle Resource for Biocomputing, Visualization, and Informatics Iliana Avila-Campillo², Hood Lab, Institute for Systems Biology, Seattle
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
